Research projects and interests
- Conformational dynamics, assembly pathways
and driving forces of spontaneous steric zipper peptide oligomerization
- Validation of empirical model parameters used in atomistic simulations
- Effect of phospholipid membranes on the self-assembly process of model peptide aggregates
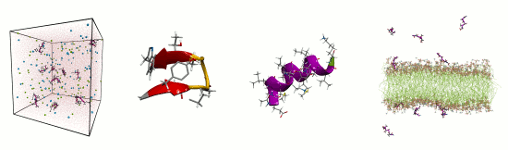
A detailed description of all the research projects currently ongoing in the Computational Biomolecular Dynamics Group can be found
here
Publications
D. Matthes, V. Gapsys, J. T. Brennecke, and B. L. de Groot,
An Atomistic View of
Amyloidogenic Self-assembly: Structure and Dynamics of Heterogeneous
Conformational States in the Pre-nucleation Phase,
Sci. Reports 2016, 33156.
D. Matthes, V. Daebel, K. Meyenberg, D. Riedel, G. Heim, U. Diederichsen, A. Lange, and B. L. de Groot,
Spontaneous Aggregation of the Insulin-Derived Steric Zipper Peptide VEALYL Results in Different Aggregation
Forms with Common Features,
J. Mol. Biol. 2014, 426,
362-376.
D. Matthes, V. Gapsys, and B. L. de Groot,
Driving forces and structural determinants of steric zipper peptide oligomer formation elucidated by atomistic simulations,
J. Mol. Biol. 2012, 421, 390-416.
D. Matthes, V. Gapsys, V. Daebel, and B. L. de Groot,
Mapping the conformational dynamics and pathways of spontaneous steric zipper peptide oligomerization,
PLoS ONE 2011, 6, e19129.
D. Matthes, and B. L. de Groot,
Secondary structure propensities in peptide folding simulations:
A systematic comparison of molecular mechanics interaction schemes,
Biophys. J. 2009, 97, 599-608.
J. Haas, E. Voehringer-Martinez, A. Boegehold, D. Matthes, U. Hensen, A. Pelah, B. Abel, and H. Grubmueller,
Primary steps of pH-dependent insulin aggregation kinetics are governed by conformational flexibility,
ChemBioChem 2009, 10, 1816-1822.
